CELLECT: contrastive embedding learning for large-scale efficient cell tracking
Mouse experiments
All mouse experiments were performed on C57BL/6J mice (Mus musculus, 6–8 weeks old), without consideration of sex, according to the animal protocols approved by the Institutional Animal Care and Use Committee of Tsinghua University. All procedures complied with the standards of the Association for Assessment and Accreditation of Laboratory Animal Care, including animal breeding and experimental manipulations. Mice were housed in standard cages with a maximum of five mice per cage, under a 12-h reverse dark–light cycle and at an ambient temperature of 72 °F and a relative humidity of ~30%. Food and water were provided ad libitum.
Model architecture
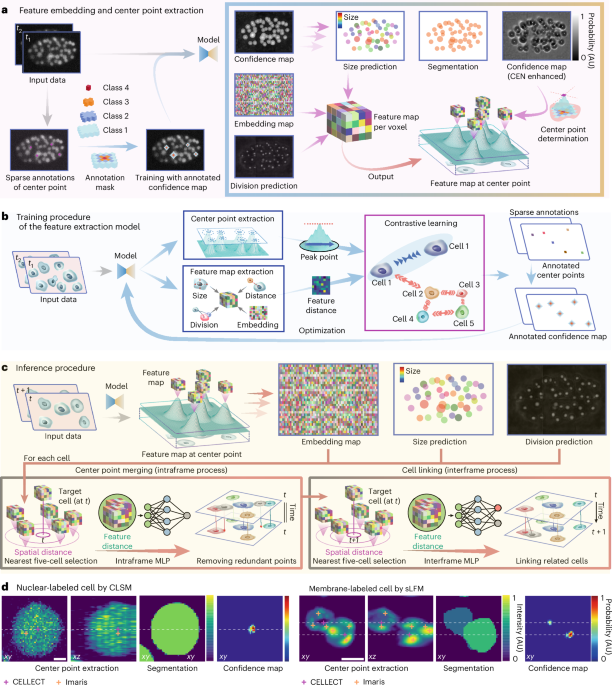
We employed a 3D U-Net architecture48 as the backbone, in which the downsampling operations use stride-2 convolutions to preserve feature granularity and spatial consistency (Supplementary Fig. 1). The primary model architecture is designed to process two consecutive imaging frames as input. The network has two output branches. The first branch outputs a confidence map for the first frame indicating the probability of being a cell center, a 64-channel latent embedding vector for each voxel and a probability map predicting whether a cell undergoes division. These outputs leverage spatiotemporal differences between consecutive frames to extract cell center-specific features and provide coarse estimates of cell size (radius), without relying on boundary-based or shape-dependent information. The second branch incorporates a lightweight 3D U-Net referred to as the CEN. The CEN sharpens the confidence maps by concentrating probability values near the cell centers, reducing the influence of intensity variations and ensuring accurate localization of cell center points. The outputs from both branches are merged to achieve precise cell center localization, segmentation and feature extraction.
To refine the tracking process, the architecture integrates two MLPs: an intraframe MLP and an interframe MLP (Supplementary Fig. 5). Each MLP takes the embedding of a target cell along with comparative features computed against five candidate cells with the nearest spatial distances as input. The obtained features, including embedding vector offsets, spatial distances, estimated size differences and changes in predicted division probabilities, were subsequently input into the intraframe and interframe MLP networks for further classification.
The intraframe MLP outputs six similarity scores: five corresponding to the candidate cells and one representing ‘none’, indicating that the target cell is not matched with any of the candidates. This ‘none’ score handles cases in which the cell disappears or moves out of the field of view. The interframe MLP not only outputs six similarity scores (five corresponding to the candidate linking cells and one representing ‘none’) but also outputs a prediction of whether the target cell is undergoing division. The detailed input–output structure of the two MLPs is shown in Supplementary Fig. 5 and Supplementary Table 3. This design enables the model to flexibly handle passive drifting of dying cells, transient cell fragments and field-of-view transitions through learned similarity scores and dynamic track assignment.
The difference between the two MLPs lies in their respective roles and operating contexts. The intraframe MLP functions within a single frame to suppress redundant or overlapping cell center predictions. It treats any two objects in the same frame as distinct cells, as long as they appear at the same time. And the MLP will attempt to remove these redundant identifications of the same cell. The interframe MLP is used to link cells between adjacent frames. It uses the division prediction to decide whether a cell at time t should be linked to one or two cells at time t + 1. In this case, daughter cells resulting from division are treated as part of the same lineage and grouped together during linking.
These two MLPs are intentionally implemented as separate modules because they rely on different input structures and perform distinct tasks. Separating them from the core feature extractor also enables efficient patch-wise inference, which is important for large-scale 3D datasets. If the MLPs were integrated into the main backbone, handling linkages across patch boundaries would become much more difficult. Compared to sequence-based models that require fixed-length temporal inputs and long sequences, this modular and lightweight design provides better generalization across varying frame intervals and temporal resolutions with reduced computational costs.
To further address the limitations of training with only sparse annotations, we adopt a spatial distance-based candidate selection strategy, leveraging previous knowledge that cells in closer proximity to the target cell are more likely to be correct linkage candidates. As shown in Supplementary Fig. 11, this spatial proximity-based approach exhibits better robustness and stability, particularly in high-density conditions or late-stage embryonic datasets with frequent mitotic events than feature distance-based selection. While feature distance-based candidate selection performs well in low-density regions, it becomes less reliable under challenging scenarios due to the sparsity of supervision in the embedding space, which often leads to incorrect associations.
Moreover, the parameter for the ‘number of candidate cells’ does not need to be fixed at five, the default parameter in CELLECT, and can be flexibly adjusted based on the frame rate and cell density of the dataset. The default configuration of selecting five candidate cells provides a good trade-off between computational efficiency and adequate coverage of potential division events. We demonstrate that five candidates are sufficient for datasets with a common frame rate (Supplementary Fig. 12). However, in data that have a low imaging frame rate and in which cells undergo larger displacements between frames, expanding the candidate pool to ten can improve matching coverage and reduce false negatives in densely populated regions.
Center enhancement network
The CEN employs a simplified and effective method for center point extraction, which directly identifies local maxima within a designated search mask zone, bypassing the complexities of traditional nonmaximum suppression49,50.
For training the center point extraction module, a 7\(\times\)7\(\times\)7 region surrounding the labeled center point is selected. The voxel at the exact center point is labeled as level 4, with adjacent voxels progressively categorized into levels 3, 2, 1 and 0 based on increasing distance. This creates five distinct levels. During training, the model learns to distinguish these levels, accumulating scores across them using the following loss function:
$$_=\mathop\limits_^\mathop\limits_{j=i}^{4}\mathrm{CrossEntropy}\left(m(a,j),n(b,i)\right)$$
$$m\left(a,j\right)=\mathrm{concat}(\max \left({a}_{0},{a}_{1},\cdots ,{a}_{j-1}\right),{a}_{j})$$
$$n\left(b,i\right)=\left\{\begin{array}{l}1\,b\ge i\\ 0\,b=0,\end{array}\right.$$
where a represents the predicted scores corresponding to the four levels from 1 to 4 and b denotes the label function, which represents the manual division into four levels. The function m(a, j) takes its maximum value in the range (0, j − 1) when calculating the score for each level; subsequently, this value is compared with aj at the jth level. The function n(b, i) is designed for selecting the computational area. Here, areas in b with a value greater than or equal to i are identified as positive samples and those with a value equal to 0 are treated as negative samples. Values not meeting either of these criteria are excluded from the loss calculation.
The choice of search mask size considerably impacts performance, as confirmed by ablation studies (Supplementary Fig. 10). Larger masks increase false negatives due to difficulty isolating individual center points, while smaller masks raise false positives by capturing multiple peaks within a cell. This balance is particularly sensitive to axial resolution differences, which influence mask effectiveness. The 5\(\times\) 5\(\times\) 3 search mask achieves the best trade-off, accommodating anisotropic resolutions while maintaining high accuracy.
The level configuration also plays a crucial role in CEN performance (Supplementary Fig. 8). Reducing the number of levels, especially higher levels such as 3 and 4, weakens the confidence gradient around the center point, resulting in multiple redundant peaks within the same cell. This increases the difficulty of linking cells across frames and leads to reduced tracking accuracy. Conversely, including all levels from 1 to 4 maintains robust confidence distributions and ensures precise center localization.
Compared to Gaussian mask and nonmaximum suppression approaches, such as those used in linajea, the CEN directly prioritizes accurate center localization without relying heavily on temporal integration. By selecting the most prominent peaks and avoiding redundant predictions, the CEN enables more efficient feature extraction and contrastive learning. This approach ensures robust performance in datasets with dense cell populations and anisotropic resolutions, making it highly effective for real-world biological imaging scenarios.
We compared structural variations of the framework: CELLECT with CEN, and CELLECT without CEN (Supplementary Fig. 7). Results indicate that incorporating the CEN improves tracking performance under the current output settings. The CELLECT-with-CEN configuration had lower false negative rates and demonstrated more stable overall performance. We further conducted a quantitative ablation study (Supplementary Table 4) demonstrating that CELLECT’s core modules, such as CEN, intraframe MLP and interframe MLP, are highly interdependent. Thus, disabling any of the modules substantially degrades tracking accuracy and stability, confirming their essential roles in reliable cell tracking.
Contrastive learning
Contrastive learning is a training approach that focuses on quantifying similarities among objects. Unlike conventional methods that require the explicit specification of several levels, this method permits category determination based on distances between feature vectors of objects. This approach has already found extensive application in tasks such as face recognition and reidentification51, which require adaptation to unknown categories.
The core principle of contrastive learning involves minimizing the discrepancies between an object’s feature vector and those of other objects belonging to the same level while maximizing differences from the feature vectors of objects belonging to different levels. In this project, we extracted a 64-dimensional embedding vector for each cell center point. By applying the triplet loss optimization methodology, we ensured that each cellular feature vector was uniquely distinguishable with respect to the Euclidean distance.
$${\mathrm{Loss}}_{\mathrm{triplet}}=\mathop{\sum }\limits_{i}^{N}{({\Vert f({{\bf{x}}}_{i})-f({{\bf{x}}}_{i}^{{\rm{p}}})\Vert }_{2}^{2}-{\Vert f({{\bf{x}}}_{i})-f({{\bf{x}}}_{i}^{{\rm{n}}})\Vert }_{2}^{2}+\alpha )}_{+},$$
where \(f\left(\bullet \right)\) refers to the model function; xi denotes the ith center point; \({{\bf{x}}}_{i}^{{\rm{p}}}\) represents a positive sample, namely, a sample from the same cell as the ith center point; and \({{\bf{x}}}_{i}^{{\rm{n}}}\) signifies a negative sample, that is, a sample from a different cell than the ith center point. The ‘+’ symbol on the lower right indicates that only values greater than zero in the equation are summed. α is a constant pertaining to the distance boundary, which was set to 0.3. In our study, we aimed to refine the feature distances both for the same cell between consecutive frames (one versus one) and for divided cells (one versus two) relative to other cells. The obtained features were subsequently input into an MLP network for further classification.
Training and inference
The training process of CELLECT is designed to refine feature representations, optimize confidence maps and classify cell relationships accurately. This modular pipeline integrates multiple objectives, ensuring robust tracking and segmentation across diverse datasets.
To achieve effective feature separation, the model employs a contrastive learning framework. The optimization process minimizes feature distances within the same cell while maximizing distances between different cells. This is formalized as:
$${E}_{{\bf{x}}\approx p({\bf{x}})}{\left(\mathop{\max }\limits_{{{\bf{y}}}^{+}\approx p({\bf{y}}{|}{\bf{x}})}\varphi ({\bf{x}},{{\bf{y}}}^{+})-\mathop{\min }\limits_{{{\bf{y}}}^{-}\approx q({\bf{y}}{|}{\bf{x}})}\varphi ({\bf{x}},{{\bf{y}}}^{-})\right)}_{+}.$$
Here, E represents the expectation, averaging the loss across all labeled sample points x, while φ(x, y) measures the similarity between the feature vectors of x and y, typically computed as the Euclidean distance. Positive samples y+ ≈ p(y|x) are defined as points originating from the same cell or dividing cells, while negative samples y− ≈ q(y,|,x) represent points from distinct labeled cells. The \(\max \left(\bullet \right)\) operator in this context selects the most distant positive sample for a given input x, ensuring that the training process focuses on maximizing the separation of the hardest-to-cluster positive pairs within the same cell. Conversely, the \(\min (\bullet )\) operator selects the nearest negative sample for x, which corresponds to the most challenging negative pair near the decision boundary. By optimizing for these specific positive and negative sample pairs, the model achieves tighter clustering within the same cell while simultaneously pushing apart distinct cells in the feature space.
For training, cross-entropy loss was employed to optimize segmentation, division probability prediction and MLP classification tasks, while the mean absolute error loss was used for cell size assessment. This combination of loss functions allowed the model to address diverse objectives simultaneously, ensuring consistent performance across tasks.
The model was trained on the mskcc-confocal dataset for 149 epochs, using the first 270 frames for training and reserving 50 frames for testing. Each epoch consisted of 400 steps, with random patches sampled from the training frames and 130 samples randomly selected from the testing frames. Input data were normalized based on the specific dataset. For datasets with fixed intensity ranges, such as nih-ls (0–255), no additional preprocessing was required. For datasets such as mskcc-confocal, which lack a fixed upper intensity bound, a logarithmic transformation was applied as input = log (input + 1).
To ensure stable optimization, MLP modules were selectively trained during specific epochs (2–7, 15–20 and 35–40), with their parameters frozen during the remaining epochs. This selective training approach minimized interference among tasks, enabling focused optimization. The Adam optimizer with a learning rate of 0.0001 was used for training, and no additional data augmentation was applied.
During inference, the model processes large images using a blockwise strategy. Images are divided into partially overlapping patches, which are processed in parallel. Using the confidence maps and search masks, cell center points are identified, retaining only the feature information corresponding to these points for downstream classification. This distributed processing strategy enables scalability and efficiency for large-scale datasets. The reproducibility of the approach was validated in five independent training runs, as shown in Supplementary Figs. 13 and 14, with consistent results demonstrating reliable, low-error tracking on the mskcc-confocal dataset.
By combining modular training with a robust optimization framework, CELLECT ensures scalable, accurate tracking and segmentation under diverse imaging conditions. The integration of contrastive learning enhances feature separation, while selective training of MLP modules and efficient inference strategies improve the overall robustness and adaptability of the model.
Temporal linking with larger intervals
CELLECT is not limited to linking cells between strictly adjacent frames. As shown in Supplementary Fig. 14, even when trained only on datasets with the original frame rate, CELLECT maintains stable tracking performance across longer intervals, such as linking from frame t to t + 2 or t + 3, in rapidly developing sequences such as C. elegans embryogenesis. While the majority of trajectories remain stable, a moderate decrease in performance is observed in late-stage sequences characterized by high cell density and frequent mitotic events. To further investigate this effect, we evaluated the impact of varying temporal downsampling during training and different settings for selecting the number of candidate cells (Supplementary Figs. 11 and 12). The results indicate that CELLECT performs reliably under common frame rate conditions using five spatial distance-based candidate cells. For lower imaging frame rate conditions, incorporating multitemporal training samples and increasing the number of spatial candidates improve robustness by reducing false negatives in crowded, actively dividing regions.
In addition to extended cross-frame inference, CELLECT supports postprocessing optimization strategies that operate on previously inferred trajectories. These include global trajectory planning, reconnection of fragmented tracks and pruning of redundant branches. For example, in the Drosophila neuron-tracking task (Fig. 5), in which signal dropout is common, we applied a ±200-frame temporal window to reconnect fragmented tracks based on feature similarity and spatial continuity. This strategy improves trajectory completeness without large computational costs. For consistency, Imaris and TrackMate were also configured with a maximum 2,000-frame temporal linking window, following the recommended settings.
These two complementary strategies, extended temporal matching during inference and postprocessing optimization based on trajectory structure, enhance CELLECT’s adaptability to a wide range of temporal resolutions and imaging conditions. It is important to note that these procedures were applied exclusively to the Drosophila neuron dataset (Fig. 5). All other results presented in this work were obtained using CELLECT’s default frame-by-frame inference without any additional temporal or postprocessing mechanisms.
Segmentation and radius estimation
While CELLECT does not rely on dense segmentation masks, it provides coarse segmentation and radius estimation based solely on sparse center point annotations. During training, approximate cell sizes were derived from radius values associated with each annotated center (for example, in the mskcc-confocal dataset) or from silver-standard masks in datasets such as Fluo-N3DH-CE. These estimates were used to define local neighborhoods for contrastive learning, where voxels within a specified radius were treated as belonging to the same cell.
The model learns to encode instance-specific spatial embeddings by minimizing intra-cell distances and maximizing inter-cell distances within local neighborhoods. This results in soft but biologically coherent segmentation maps around center points, even in the absence of boundary-level supervision. Although CELLECT does not explicitly optimize precise boundary delineation, the resulting coarse segmentation remains highly informative for downstream tasks such as cell–cell interaction analysis, as demonstrated in Fig. 4.
Supplementary Fig. 3 further illustrates the smooth clustering of embedding vectors around the center points, and Supplementary Fig. 4 shows how spatial convolution of the embedding map helps delineate cellular boundaries in an unsupervised fashion. We note that the grid-patterned artifacts observed in Supplementary Fig. 4 result from the independent inference performed on each patch. However, these artifacts do not impact performance, as only the embedding vectors at the cell centers are used.
Comparison with embedding-based and contrastive learning methods
CELLECT shares conceptual similarities with previous instance segmentation frameworks that use learned embeddings to distinguish object instances, such as that of Neven et al.52,53. Their model predicts pixel-wise spatial embeddings and applies clustering as a postprocessing step to derive object masks. However, this approach requires dense, per-pixel supervision and is designed for natural images with full segmentation annotations. By contrast, CELLECT is tailored for sparse annotation settings typical of 3D microscopy, in which only center points are available. Our architecture directly learns object-level representations from these sparse annotations without relying on full-resolution masks or clustering. In short, Neven’s model learns centers from full images; CELLECT learns object structure from sparse centers.
We also compare CELLECT with the recent contrastive learning approach of Zyss et al.28, which focuses on lineage reconstruction from fully segmented cells. Their framework is optimized for low-imaging-frame rate datasets lacking trajectory labels and infers division events from appearance-based cell similarity. CELLECT, by comparison, is designed for high-resolution 3D time-lapse microscopy, for which accurate cell identity tracking is required in dynamic environments with frequent mitosis. While both methods leverage contrastive learning, Zyss et al. aim to recover lineage trees, whereas CELLECT emphasizes short-term identity association and frame-to-frame correspondence.
In sum, these methods differ from CELLECT in annotation regime (dense masks versus sparse centers), learning objective (lineage inference or instance segmentation versus tracking) and data modality (two-dimensional natural or static bioimages versus dynamic 3D microscopy). We view these approaches as complementary: each addressing different challenges in the broader problem of cell analysis.
Evaluation with the Cell Tracking Challenge benchmark
To evaluate CELLECT’s generalizability on a standardized third-party test set, we submitted our method to the Cell Tracking Challenge under the team named ‘THU-CN (3)’. Please note that the number (3) in the name THU-CN means that there are two earlier submissions under the same institution that were actually submitted several years ago by other research teams at Tsinghua University and not our group. On the Fluo-N3DH-CE dataset, CELLECT achieved overall performance scores of 0.853 in the segmentation benchmark and 0.850 in the tracking benchmark, indicating the highest performance in both tasks.
We selected the Fluo-N3DH-CE dataset because it is currently the most suitable 3D time-lapse dataset in the Cell Tracking Challenge that provides complete training annotations, including cell center positions, trajectories and mitosis events. Its annotation structure and biological context align well with our training design, and it stands out as a rare 3D dataset within the Cell Tracking Challenge that supports comprehensive and integrated evaluation across segmentation, mitotic event detection and tracking metrics.
While the official training subset of Fluo-N3DH-CE was used to prepare the Cell Tracking Challenge submission, this dataset was not involved in any training or evaluation experiments described in the main text. All results were independently evaluated by Cell Tracking Challenge organizers using hidden ground truth annotations. The full results are available at and https://celltrackingchallenge.net/latest-ctb-results/.
Comparative methods and parameter settings
To ensure fair and reproducible benchmarking, we evaluated CELLECT against representative and widely used tracking methods. For Imaris (Bitplane, version 9.0.1), we used the ‘Spots’ detection and ‘Tracking’ modules. Estimated spot diameters were manually adjusted based on voxel size for each dataset. Gap closing was enabled with a maximum of two to three frames for standard datasets and extended to 2,000 frames for neuronal sequences to accommodate intensity flickering. For TrackMate54 (via ImageJ, version 7.10.1), we used the provided pretrained Weka55 model detector applied in a slice-by-slice manner. The gap-closing interval was set to 2,000 frames for neuronal recordings, consistent with the Imaris configuration. For StarryNite34,36,37 (2020 release), we adopted the default parameter configuration distributed with the 2020 version of StarryNite and modified the cell radius range settings based on the ground truth annotation files.
C. elegans embryo public dataset
The data used in the quantitative benchmarking of CELLECT were obtained from two public imaging datasets of C. elegans embryo development, acquired using a confocal microscope (mskcc-confocal, and a light sheet microscope (nih-ls, respectively. The confocal dataset contains three anisotropic 3D time-lapse series, while the light sheet dataset contains three isotropic 3D time-lapse series, both featuring sparse cell annotations for all cells. These datasets were employed to evaluate the performance metrics of CELLECT based on the provided ground truth. Each dataset contains three sequences, and each sequence comprises a single embryo imaged over 400 frames with sparse center point annotations.
We adopted a threefold cross-validation protocol following the linajea framework. Specifically, for each dataset, the first 270 frames of each sequence were used. In each round, two sequences were used for training and validation and the remaining one was reserved for testing. This setup ensures strict separation between training and testing data and allows fair comparison with previous work. All other experiments in the study also used only the first 270 frames from each sequence. The results presented in Fig. 2 were obtained using this cross-validation protocol. Detailed evaluation results for each fold are reported in Supplementary Tables 1 and 2.
Dual-channel T cell imaging in lymph nodes
To achieve simultaneous sparse and dense labeling of the same T cells across different fluorescence imaging channels, we used transgenic mice coexpressing OT-II DsRed and OT-II GFP at a ratio of 1:5. The sparsely labeled DsRed channel, tracked using Imaris, was used as the gold standard to evaluate tracking performance of the same cells in the densely labeled GFP channel. We used a two-photon microscope to acquire dual-channel imaging datasets at 30 Hz per plane across 50 planes. For an improved signal-to-noise ratio, results from every 20 frames were averaged, resulting in a final frame interval of 33.33 s and a 2.5-h 3D time-lapse series dataset. The imaging area covered 688 μm × 688 μm × 100 μm.
B cell imaging during germinal center formation
To observe the active behaviors of B cell during GC formation, a surgical procedure for intravital imaging of the inguinal lymph node was performed. Mice were anesthetized with a mixture of isoflurane and oxygen, the surgical area was shaved, and residual hair was removed with depilatory cream. Following established procedures56,57, a stable imaging window of the lymph node was prepared. The mouse was then securely fixed under the stage of a 2pSAM instrument for imaging17. Intravital imaging was performed over 12 h with a frame interval of 33.90 s per two planes, with each plane covering a thickness of 100 μm. The image area covered 688 μm × 688 μm × 200 μm.
Imaging of tripartite interactions among bacteria, neutrophils and macrophages
To investigate tripartite interactions among bacteria, neutrophils and macrophages, we used Streptococcus pneumoniae (TH870, serotype 6A) expressing GFP, neutrophils labeled with PE-Cy5-conjugated anti-Ly6G antibody and splenic red pulp macrophages labeled with PE-conjugated anti-F4/80 antibody. For intravital imaging, the spleen was exposed, stabilized using an adsorption pump and observed through an 8-mm coverglass window under anesthesia using 2pAM. Thirty minutes after tail vein injection of 107 CFU of GFP-expressing bacteria, 3D imaging was performed at 30 volumes per second, covering the volume of 229 μm × 229 μm × 25 μm.
Drosophila brain imaging under tissue deformation
Flies expressing pan-neuronal GCaMP6s (nSyb-Gal4, UAS-nls-GCaMP6s) were raised on standard cornmeal medium with a 12-h light and 12-h dark cycle at 23 °C and 60% humidity and housed in mixed male–female vials. Female flies (3–8 d old) were selected for brain imaging. UAS-nls-GCaMP6s flies were provided by D.J. Anderson (California Institute of Technology). To prepare for imaging, flies were anesthetized on ice and mounted in a 3D-printed plastic disk. The posterior head cuticle was opened using sharp forceps (5SF, Dumont) at room temperature in fresh saline (103 mM NaCl, 3 mM KCl, 5 mM TES, 1.5 mM CaCl2, 4 mM MgCl2, 26 mM NaHCO3, 1 mM NaH2PO4, 8 mM trehalose and 10 mM glucose (pH 7.2), bubbled with 95% O2 and 5% CO2). After this, the fat body and air sac were also removed carefully. The position and angle of the flies were adjusted to keep the posterior of the head horizontal, and the window was made large and clean, for the convenience of observing multiple brain regions. Brain movement was minimized by adding UV glue around the proboscis58. Due to motion and latent osmotic pressure mismatch between the imaging saline and the body fluid of the fly, the brain showed deformation and nucleus drift during the imaging period.
We used a 2pSAM (mid-NA) system implemented with a water-immersion objective lens (×25, 1.05 NA, Olympus). Intravital volume imaging was acquired at a frame rate of 30 Hz with 13-angle scanning. The lateral field of view, 229 μm × 229 μm, covered the entire lateral range of the left central brain and some parts of the optic lobe. For continuous imaging over about 2 h, a wavelength of 920 nm at 25–35 mW of power was used for excitation. For preprocessing and reconstruction, we applied DeepCAD-RT to the images of each angle to enhance the signal-to-noise ratio59,60. Customized denoising models were trained for each channel of each fly. Next, we reconstructed the volumes using the algorithm of 2pSAM17. After the reconstruction, the lateral and axial resolutions were 0.45 μm and 0.75 μm, respectively.
Data analysis
All data processing and analysis were performed using Python (version 3.9). The temporal traces of neurons in Fig. 5 were computed using the formula ΔF/F0 = (F − F0)/F0, where F represents the intensity at the center points of neurons and F0 denotes the mean value of F. Imaris (version 9.0.1) and TrackMate (version 7.10.1 in ImageJ) were used for cell tracking and segmentation analysis.
Statistics and reproducibility
No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment. Figure 2 is based on public datasets with cross-validation and does not involve independent experimental replication. Figures 3–5 are based on existing microscopy datasets, and the experiments were performed once without independent replication.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link








